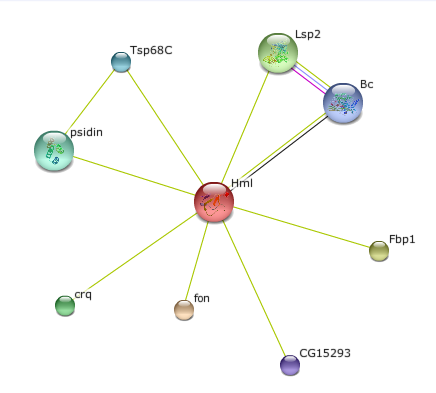
Protein Interactions of the Hemolectin protein in Drosophila melanogaster
Bc and Lsp2 proteins
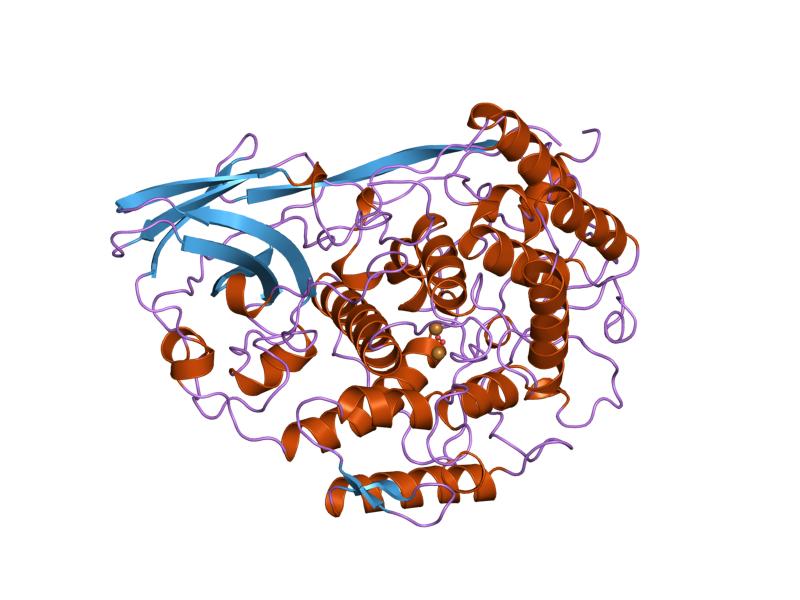
Bc GO: Oxygen transporter activity, response to wounding, scab formation [4]
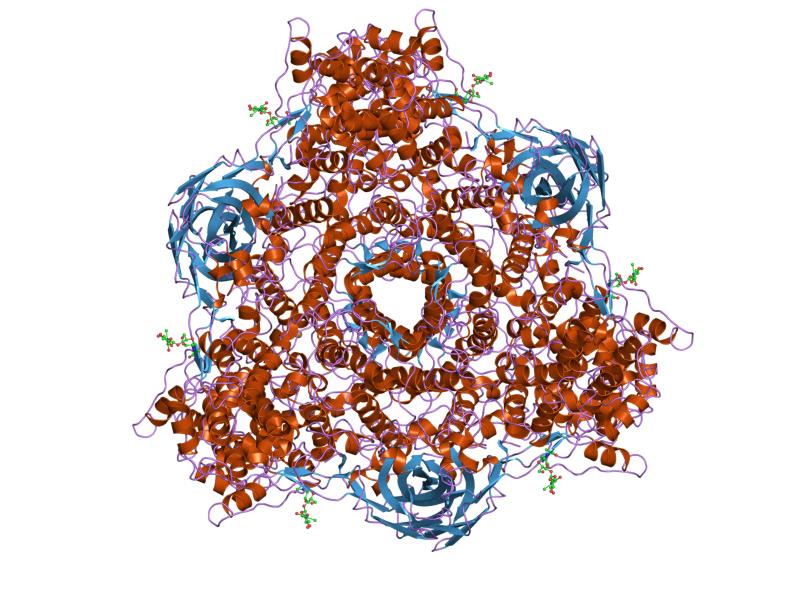
Lsp2 GO: Extracellular region, larval serum protein complex, oxygen transporter activity [5] Larval serum protein 2; Larval storage protein (LSP) which may serve as a store of amino acids for synthesis of adult proteins [string-db]
Lsp2 GO: Extracellular region, larval serum protein complex, oxygen transporter activity [5] Larval serum protein 2; Larval storage protein (LSP) which may serve as a store of amino acids for synthesis of adult proteins [string-db]
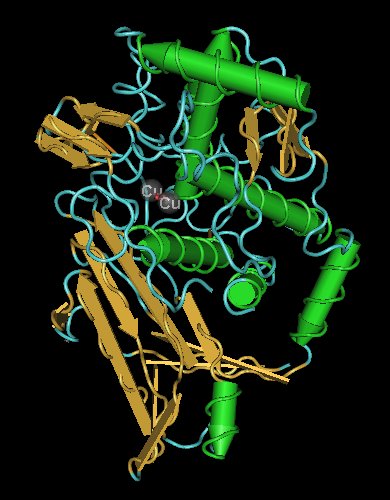
Hemocyanins are proteins that transport oxygen throughout the bodies of some invertebrate animals. These metalloproteins contain two copper atoms that reversibly bind a single oxygen molecule (O2). They are second only to hemoglobin in frequency of use as an oxygen transport molecule. Unlike the hemoglobin in red blood cells found invertebrates, hemocyanins are not bound to blood cells but are instead suspended directly in the hemolymph. Oxygenation causes a color change between the colorless Cu(I) deoxygenated form and the blue Cu(II) oxygenated form.
http://pfam.sanger.ac.uk/family/PF03722.9
http://pfam.sanger.ac.uk/family/PF03722.9
Tsp68C
GO: Integral to membrane, hemopoiesis [6]
|
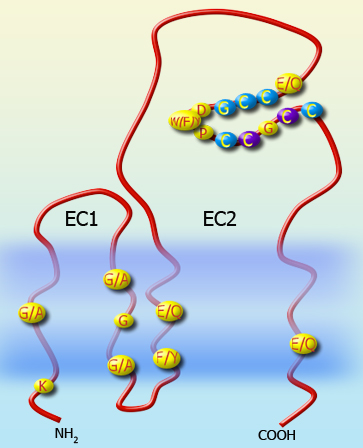
Tetraspanins, also called tetraspans or the transmembrane 4 superfamily (TM4SF), have four transmembrane domains, intracellular N- and C-termini and two extracellular domains, one short (called the small extracellular domain or loop, SED/SEL or EC1) and one longer, typically 100 amino acid residues (the large extracellular domain/loop, LED/LEL or EC2). Although several protein families have four transmembrane domains, tetraspanins are defined by conserved domains listed in the Protein Families database under pfam00335.12.[1] The key features are four or more cysteine residues in the EC2 domain, with two in a highly conserved 'CCG' motif.
Research into this field is relatively recent (less than 20 years) and therefore there is much to learn about the function of specific tetraspanins. Generally, tetraspanins are often thought to act as scaffolding proteins, anchoring multiple proteins to one area of the cell membrane.[2] Tetraspanins are highly conserved between species. Some tetraspanins can have N-linked glycosylations on the long extracellular loop (LEL, EC2) and palmitoylations at a CXXC motif in their transmembrane region.[3] There are 34 tetraspanins in mammals, 33 of which have also been identified in humans. Tetraspanins display numerous properties that indicate their physiological importance in cell adhesion, motility, activation and proliferation, as well as their contribution to pathological conditions such as metastasis or viral infection. A role for tetraspanins in platelets was demonstrated by the bleeding phenotypes of CD151- and TSSC6-deficient mice, which exhibit impaired "outside-in" signalling through αIIbβ3, the major platelet integrin. it is hypothesized that tetraspanins interact with and regulate other platelet receptors.[4] http://pfam.sanger.ac.uk//family/PF00335.15 |
Psidin
GO: Lysosome, binding, humoral immune response, phagocytosis [7] phagocyte signaling impaired; Has 2 roles in the larval immune response: required both for the phagocytic degradation of internalized bacteria and for the induction of Defensin in the fat body. Within the phagocytic blood cells, has a role in detection of infection and activation of the humoral immune response [string-db]
N-acetyltransferase B complex (NatB) non catalytic subunit
This is the non-catalytic subunit of the N-terminal acetyltransferase B complex (NatB). The NatB complex catalyses the acetylation of the amino-terminal methionine residue of all proteins beginning with Met-Asp or Met-Glu and of some proteins beginning with Met-Asn or Met-Met. In Saccharomyces cerevisiae this subunit is called MDM20 and in Schizosaccharomyces pombe it is called Arm1. NatB acetylates the Tpm1 protein and regulates and tropomyocin-actin interactions. This subunit is required by the NatB complex for the N-terminal acetylation of Tpm1 [1].
http://pfam.sanger.ac.uk//family/PF09797.4
This is the non-catalytic subunit of the N-terminal acetyltransferase B complex (NatB). The NatB complex catalyses the acetylation of the amino-terminal methionine residue of all proteins beginning with Met-Asp or Met-Glu and of some proteins beginning with Met-Asn or Met-Met. In Saccharomyces cerevisiae this subunit is called MDM20 and in Schizosaccharomyces pombe it is called Arm1. NatB acetylates the Tpm1 protein and regulates and tropomyocin-actin interactions. This subunit is required by the NatB complex for the N-terminal acetylation of Tpm1 [1].
http://pfam.sanger.ac.uk//family/PF09797.4
Crq
GO: Cell adhesion, immune response, phagocytosis [3] Macrophage receptor for apoptotic cells [string-db]
|
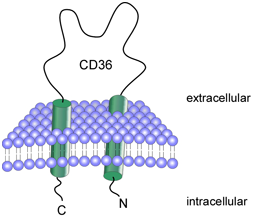
CD36 (Cluster of Differentiation 36) is an integral membrane protein found on the surface of many cell types in vertebrate animals and is also known as FAT, SCARB3, GP88, glycoprotein IV (gpIV) and glycoprotein IIIb (gpIIIb). CD36 is a member of the class B scavenger receptor family of cell surface proteins. CD36 binds manyligands including collagen,[1] thrombospondin,[2] erythrocytes parasitized with Plasmodium falciparum,[3] oxidizedlow density lipoprotein,[4][5] native lipoproteins,[6] oxidized phospholipids,[7] and long-chain fatty acids.[8]
Recent work using genetically modified rodents have identified a clear role for CD36 in fatty acid and glucosemetabolism,[9][10] heart disease,[11] taste,[12] and dietary fat processing in the intestine.[13] It may be involved in glucose intolerance, atherosclerosis, arterial hypertension, diabetes, cardiomyopathy and Alzheimer's disease.[14] |
Fon
|
Gene ontology:
Hemolymph coagulation and Metamorphosis [1] |
CG15293
Fbp1
GO: Oxygen transport activity, Protein transport activity, storage protein import into fat body. [2]
Sources
http://string-db.org/
1 http://refgene.com/gene/35211
2 http://refgene.com/gene/39566
3 http://refgene.com/gene/33219
4 http://refgene.com/gene/37044
5 http://refgene.com/gene/45326
6 http://refgene.com/gene/117407
7 http://refgene.com/gene/42389
1 http://refgene.com/gene/35211
2 http://refgene.com/gene/39566
3 http://refgene.com/gene/33219
4 http://refgene.com/gene/37044
5 http://refgene.com/gene/45326
6 http://refgene.com/gene/117407
7 http://refgene.com/gene/42389